This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein domains?
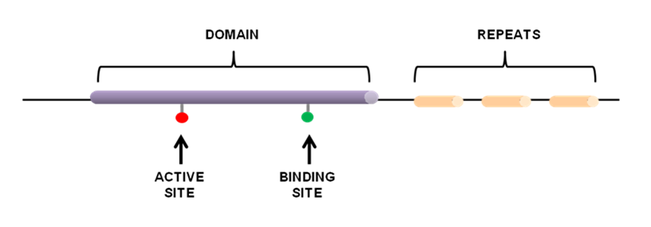
Protein domains are distinct functional and structural units found in amino acid sequences. These domains usually have specific functions or interactions and a single protein can contain multiple functional sites. Protein domains often consist of sequence features. Sequence features are small groups of amino acids that are only a few amino acids long and have characteristics important for the function of a protein [1]. Examples of sequence features are:
- active sites: involved in catalytic activity such as in enzymes.
- binding sites: contain amino acids involved in binding molecules.
- post translational modification sites: residues known to be chemically modified via phosphorylation, palmitoylation, acetylation and etc.
- repeats: short sequences that are repeated and have specific binding or structural properties essential for protein function.
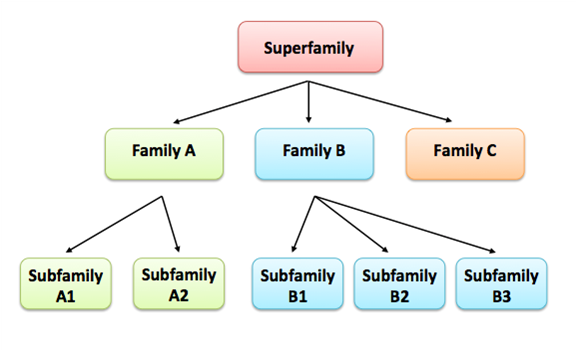
Proteins also are arranged in hierarchical order, with proteins sharing a common ancestor being subdivided into smaller and more closely related groups. The order (superfamily, family and subfamily) is depicted below.
What protein domains are found in MEF2C?
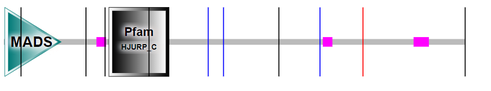
SMART, PFAM, InterPRO, and PROSITE were used to search for protein domains in MEF2C. Most sources yielded different results with one domain being the only constant in the searches. The results obtained are listed below.
In all sources, the MADS box transcription factor (named after MCM1, AG, DEFA and SRF) is the common domain found in MEF2C. PFAM showed a SRF-like transcription factor in the first site whereas InterPRO showed a MEF2 (myocyte enhancer factor 2)-like transcription factor in the first site. MEF2-like transcription factor is a subfamily of the MADS superfamily whereas SRF-like transcription factor is a Type-I subfamily of the MADS superfamily. SMART, PFAM and InterPRO also had Holliday junction regulator protein family C-terminal (HJURP_C) as the second domain site in MEF2C. Hence, the main two protein domains found in MEF2C is the MADS box transcription factor and HJURP_C.
In addition to these sites, literature review shows that there are important post translational modification sites in MEF2C which are phosphorylation sites required in neuronal development in humans [2]. These sites are serine at position 387 (S387), threonine at position 293 (T293) and threonine at position 300 (T300), with S387 being the most important site [2]. A more in depth discussion of phosphorylation in the role of MEF2C can be found in the post-translational modifications page.
In addition to these sites, literature review shows that there are important post translational modification sites in MEF2C which are phosphorylation sites required in neuronal development in humans [2]. These sites are serine at position 387 (S387), threonine at position 293 (T293) and threonine at position 300 (T300), with S387 being the most important site [2]. A more in depth discussion of phosphorylation in the role of MEF2C can be found in the post-translational modifications page.
Homology of domains
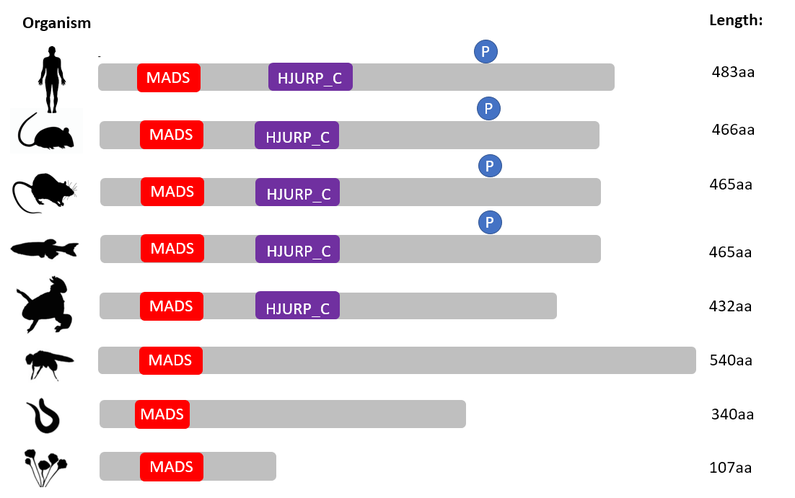
Discussion:
Since MADS is a protein superfamily, it is not surprising that this protein is conserved across all species. The HJURP_C domain is conserved mainly in vertebrates because it is a vertebral protein, hence the percent identities of MEF2C is higher in vertebrates compared to invertebrates [3]. Another important aspect of the function of MEF2C is its phosphorylation sites. Phosphorylation of the S387 site as well as other sites are required for transcriptional activation of MEF2C by Extracellular signal-regulated kinase (ERK5) and p38 mitogen-activated protein kinase (p38 MAPK). Phosphorylation of these sites are important in the survival of developing neurons. Hence, this will be investigated more in depth in the specific aims section.
References:
[1] https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-sequence-features
[2] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
[3] www.ebi.ac.uk/interpro/entry/IPR021052
Images:
Header: http://wozniaklab.com/pumilio_rna.html
Fig. 1 & 2: [1]
[1] https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-sequence-features
[2] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
[3] www.ebi.ac.uk/interpro/entry/IPR021052
Images:
Header: http://wozniaklab.com/pumilio_rna.html
Fig. 1 & 2: [1]