This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Gene Ontology?
With the advent of bioinformatics and genomics, more data is being produced and scientists need a way to systematically categorize the functions and locations of gene products. Gene ontology is a way of using standardized terms to categorize gene functions (GO terms) and how these genes interact with each other (relations). The three ways that genes can be categorized are molecular function, cellular component and biological process. [1]
Molecular Function
Describes the function of the gene products at the molecular level such as ‘binding activity’ or ‘catalytic activity’.
Cellular Component
Describes the location in a cell where the gene product is usually found. Locations of gene products can be described as relative to cellular structures or as stable macromolecular complexes.
Biological Process
Describes series of events consisting of an organized set of molecular functions. An example is ‘oxidative phosphorylation’.
Describes the function of the gene products at the molecular level such as ‘binding activity’ or ‘catalytic activity’.
Cellular Component
Describes the location in a cell where the gene product is usually found. Locations of gene products can be described as relative to cellular structures or as stable macromolecular complexes.
Biological Process
Describes series of events consisting of an organized set of molecular functions. An example is ‘oxidative phosphorylation’.
What are the GO terms for MEF2C?
Molecular Function
MEF2C is a transcription factor and it is annotated as being involved in transcription factor activity, RNA polymerase II core promoter sequence-specific DNA binding (GO:0000983). MEF2C interacts selectively and non-covalently with specific DNA sequence in an RNA polymerase II (Pol II) core promoter to modulate transcription by Pol II [2]. It is a critical transcriptional regulator functioning at the nexus of numerous synapse and neurodevelopmental disorder-linked genes [5].
MEF2C is a transcription factor and it is annotated as being involved in transcription factor activity, RNA polymerase II core promoter sequence-specific DNA binding (GO:0000983). MEF2C interacts selectively and non-covalently with specific DNA sequence in an RNA polymerase II (Pol II) core promoter to modulate transcription by Pol II [2]. It is a critical transcriptional regulator functioning at the nexus of numerous synapse and neurodevelopmental disorder-linked genes [5].
Cellular Component
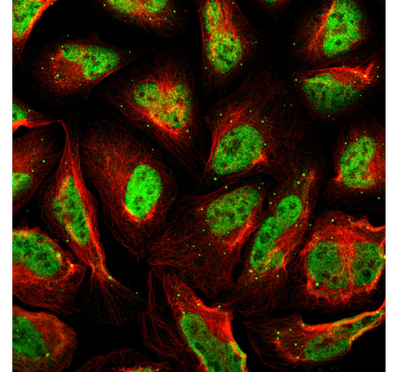
MEF2C is mainly found in the nucleus (GO:0005634) in most cells. The nucleus is a membrane-bounded organelle of eukaryotic cells in which chromosomes are located and replicated. This makes sense since it is a transcription factor and regulates gene transcription.
MEF2C is mainly found in the nucleus (GO:0005634) in most cells. The nucleus is a membrane-bounded organelle of eukaryotic cells in which chromosomes are located and replicated. This makes sense since it is a transcription factor and regulates gene transcription.
Biological process
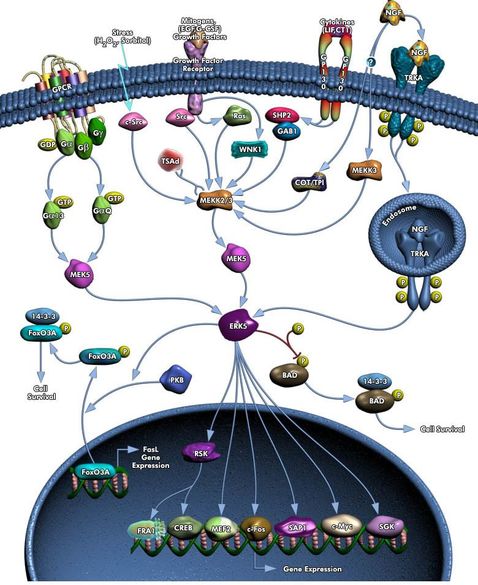
MEF2C is annotated as part of many different biological processes since it is a protein superfamily. Nevertheless, the most relevant biological processes in studying depression is its annotation in the MAPK cascade (GO:0000165) , neuron migration (GO:0001764) and neuron differentiation (GO:0030182) [2]. MEF2C is transcriptionally activated by phosphorylation by Extracellular-signal-regulated kinase 5 (ERK 5), which is a protein kinase of the MAPK family [3]. ERK 5 translocates to cell nucleus in response to extracelluar signals, where it regulates MEF2C gene expression through phosphorylation [2,4]. An extracelullar signal that influences ERK5 pathway in activation of MEF2C is neurotrophin, which is involved in survival of signalling pathways in developing neurons [3].
MEF2C is annotated as part of many different biological processes since it is a protein superfamily. Nevertheless, the most relevant biological processes in studying depression is its annotation in the MAPK cascade (GO:0000165) , neuron migration (GO:0001764) and neuron differentiation (GO:0030182) [2]. MEF2C is transcriptionally activated by phosphorylation by Extracellular-signal-regulated kinase 5 (ERK 5), which is a protein kinase of the MAPK family [3]. ERK 5 translocates to cell nucleus in response to extracelluar signals, where it regulates MEF2C gene expression through phosphorylation [2,4]. An extracelullar signal that influences ERK5 pathway in activation of MEF2C is neurotrophin, which is involved in survival of signalling pathways in developing neurons [3].
Discussion
As expected, MEF2C is annotated as a transcription factor involved in various forms of neuronal development. Despite it being annotated as a transcription factor, the specific transcriptional regulation it is responsible for has been debated. In addition, although it is identified as a transcriptional activator, more recent publications have shown that it might be a transcriptional repressor as well [3,5]. Hence, my specific aims seek to understand how MEF2C acts as a transcription factor in regulating synaptic density during brain development.
References:
[1] http://geneontology.org/page/ontology-documentation
[2] http://www.geneontology.org
[3] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
[4] https://www.phosphosite.org/
[5] Harrington, A. J., Raissi, A., Rajkovich, K., Berto, S., Kumar, J., Molinaro, G., ... & Huber, K. M. (2016). MEF2C regulates cortical inhibitory and excitatory synapses and behaviors relevant to neurodevelopmental disorders. Elife, 5.
Images:
Header: http://cissrm.ru/ceminar-texnologii-raboty-fondov-mestnyx-soobshhestv-v-regionax-rossii/
Fig.2: https://www.proteinatlas.org/ENSG00000081189-MEF2C/cell#human
[1] http://geneontology.org/page/ontology-documentation
[2] http://www.geneontology.org
[3] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
[4] https://www.phosphosite.org/
[5] Harrington, A. J., Raissi, A., Rajkovich, K., Berto, S., Kumar, J., Molinaro, G., ... & Huber, K. M. (2016). MEF2C regulates cortical inhibitory and excitatory synapses and behaviors relevant to neurodevelopmental disorders. Elife, 5.
Images:
Header: http://cissrm.ru/ceminar-texnologii-raboty-fondov-mestnyx-soobshhestv-v-regionax-rossii/
Fig.2: https://www.proteinatlas.org/ENSG00000081189-MEF2C/cell#human