This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are post-translational modifications?
Post-translational modifications (PTMs) of proteins are chemical changes made to proteins after translation to diversify the function of the proteome. These modifications are often catalyzed by specific proteins or enzymes. PTMs play an important role in the folding of proteins, their targeting to precise subcellular locations and other proteins as well as their functional state. Hence, studying PTMs via proteomics will allow us to understand how proteins specifically function [1].
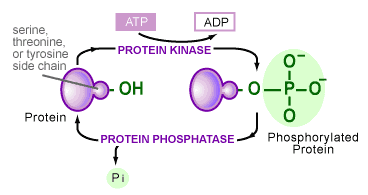
One form of posttranslational modification is phosphorylation. Phosphorylation causes conformational changes in the protein’s structure, which mediates protein-protein interactions, protein localization, protein activity and signal transduction. Phosphorylation involves attaching a phosphate group to protein residues by enzymes called kinases whereas dephosphorylation is done by phosphatases (Fig.1). The most frequently phosphorylated amino acids are serine and threionine. [2]
One form of posttranslational modification is phosphorylation. Phosphorylation causes conformational changes in the protein’s structure, which mediates protein-protein interactions, protein localization, protein activity and signal transduction. Phosphorylation involves attaching a phosphate group to protein residues by enzymes called kinases whereas dephosphorylation is done by phosphatases (Fig.1). The most frequently phosphorylated amino acids are serine and threionine. [2]
Phosphorylation of MEF2C
Discussion
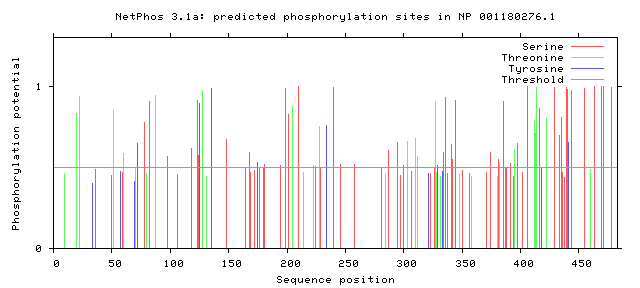
MEF2C is regulated by the Mitogen-activated protein kinase (MAPK) pathway via phosphorylation. Two major kinases that are involved in MAPK regulation of neuron development in the nervous system are p38MAPK and MAPK7 (ERK5). It has been shown that serine at position 387 (S387), threonine at position 293 (T293) and threonine at position 300 (T300) are important MEF2C phosphorylation sites, with S387 being the most important site [2]. Nevertheless, NetPhos 3.0 shows the presence of additional phosphorylation sites in human MEF2C protein but not those mentioned above. I will investigate the function of MEF2C phosphorylation sites in more detail to address this discrepancy by investigating all conserved phosphorylation sites as mentioned in my specific aims. By doing so, I will be able to check if the phosphorylation sites cited in literature are important in neuron development in addition to discovering new phosphorylation sites involved in synaptic density regulation.
References:
[1] Bürkle, A. (2001). Encyclopedia of Genetics. Posttranslational Modification.
[2] Saraswathy, N., & Ramalingam, P. (2011). Concepts and Techniques in Genomics and Proteomics. Elsevier.
[3] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
Images:
Fig.1: https://www.scq.ubc.ca/protein-phosphorylation-a-global-regulator-of-cellular-activity/
Fig.2: www.cbs.dtu.dk/services/NetPhos/
[1] Bürkle, A. (2001). Encyclopedia of Genetics. Posttranslational Modification.
[2] Saraswathy, N., & Ramalingam, P. (2011). Concepts and Techniques in Genomics and Proteomics. Elsevier.
[3] Wang, Y., Liu, L., & Xia, Z. (2007). Brain‐derived neurotrophic factor stimulates the transcriptional and neuroprotective activity of myocyte‐enhancer factor 2C through an ERK1/2‐RSK2 signaling cascade. Journal of neurochemistry, 102(3), 957-966.
Images:
Fig.1: https://www.scq.ubc.ca/protein-phosphorylation-a-global-regulator-of-cellular-activity/
Fig.2: www.cbs.dtu.dk/services/NetPhos/