This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein interaction networks?
Protein interactions connect cell components and cell processes to form protein complexes and mediate post translational modifications. Protein interaction networks represent the union of all proteins and the interactions among them at the cellular level. It allows us to map interactions between proteins, understand which properties characterize them. Most importantly, protein interaction networks allow us to understand how the relationship of protein functions to other regulatory networks such as gene regulatory networks and how they result in biological function. [1] Databases such as STRING, IntAct and Biogrid can be used to understand protein interaction networks.
A network comprises of nodes and edges. Nodes represent proteins and can be filled or unfilled. Unfilled nodes contain proteins whereby the protein structure is unknown whereas filled nodes contain proteins of known structure. Egdes show the protein-protein interactions and does not mean that proteins are physically bound to each other. [1]
A network comprises of nodes and edges. Nodes represent proteins and can be filled or unfilled. Unfilled nodes contain proteins whereby the protein structure is unknown whereas filled nodes contain proteins of known structure. Egdes show the protein-protein interactions and does not mean that proteins are physically bound to each other. [1]
MEF2C protein interaction networks
Discussion
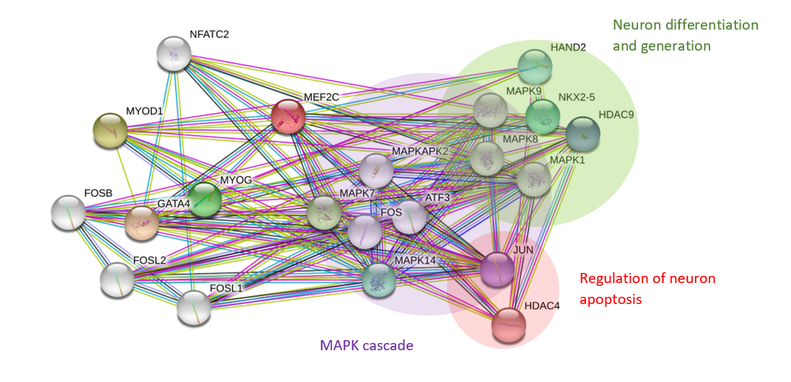
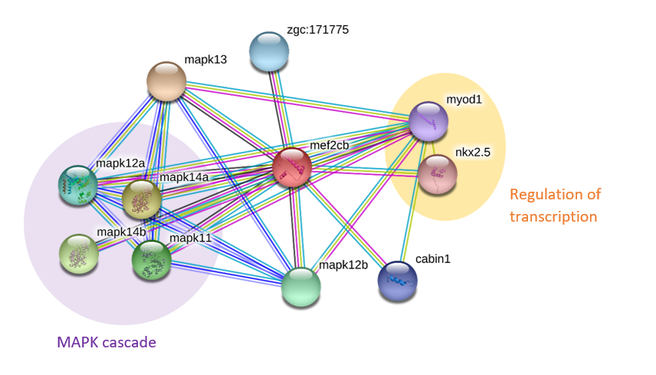
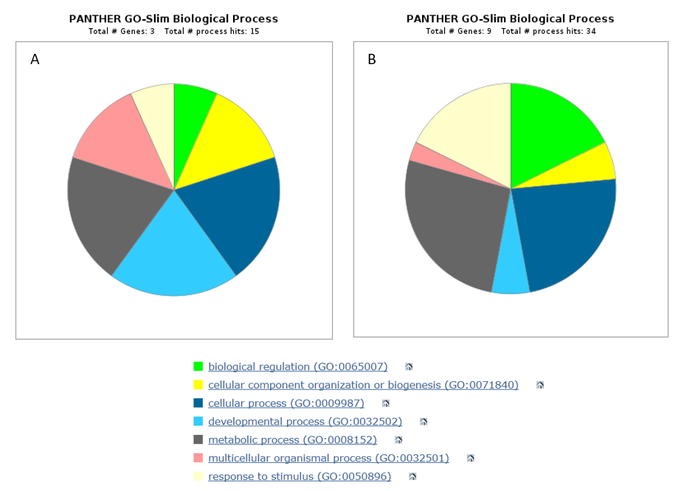
The protein interaction network of MEF2C in humans shows that interacting proteins can be classified into three major biological processes related to neuronal growth. These are neuron development and differentiation, regulation of neuron apoptosis and the MAPK cascade. Not surprisingly the network shows the presence of proteins that are involved in neuronal apoptosis, supporting earlier findings that MEF2C regulates neuronal development during maturation via apoptosis of unused neurons [2]. There are also a large number of proteins involved in the MAPK cascade, showing that the role of phosphorylation of MEF2C in promoting neuronal development is important. There are less proteins sorted by Gene Ontology terms in zebrafish but there are also a relatively large number of proteins interactors involved in the MAPK cascade. Once again, this shows that phosphorylation is an important process related to MEF2C function. PANTHER analysis of GO terms showed that most protein interactors are involved in cellular, metabolic and developmental processes and this is mostly consistent in both species.
Hence, it is clear that the MAPK cascades play a large role in MEF2C interaction and function in both humans and zebrafish. Despite a limited amount of information on how MEF2CB regulates neuronal growth in zebrafish, these MAPK cascades show promising steps forward in understanding how MEF2C regulates neuronal development and differentiation.
Hence, it is clear that the MAPK cascades play a large role in MEF2C interaction and function in both humans and zebrafish. Despite a limited amount of information on how MEF2CB regulates neuronal growth in zebrafish, these MAPK cascades show promising steps forward in understanding how MEF2C regulates neuronal development and differentiation.
Reference:
[1] https://www.ncbi.nlm.nih.gov/books/NBK56024/
[2] Barbosa, A. C., Kim, M. S., Ertunc, M., Adachi, M., Nelson, E. D., McAnally, J., ... & Olson, E. N. (2008). MEF2C, a transcription factor that facilitates learning and memory by negative regulation of synapse numbers and function. Proceedings of the National Academy of Sciences, 105(27), 9391-9396.
Images:
Header: https://towardsdatascience.com/a-soft-introduction-to-neural-networks-6986b5e3a127
[1] https://www.ncbi.nlm.nih.gov/books/NBK56024/
[2] Barbosa, A. C., Kim, M. S., Ertunc, M., Adachi, M., Nelson, E. D., McAnally, J., ... & Olson, E. N. (2008). MEF2C, a transcription factor that facilitates learning and memory by negative regulation of synapse numbers and function. Proceedings of the National Academy of Sciences, 105(27), 9391-9396.
Images:
Header: https://towardsdatascience.com/a-soft-introduction-to-neural-networks-6986b5e3a127